About
Welcome to the official website for the New York meetup of the Learning on Graphs Conference, an annual research conference that covers areas broadly related to machine learning on graphs and geometry.
This event serves as a local branch of the main conference, providing an environment for researchers in this field to convene and foster discussion and social connections.
The main objective of this event is to build an open and diverse community in the greater NYC area with students, professors, and industry researchers interested in machine learning and graphs (e.g., computer science, discrete math, operations research, etc.), geometry (e.g., applied math, physics, neuroscience, biology, etc.), and networks (e.g., network science, social science, etc.).
Explore last year event's recorded sessions here.
Learning Meets Geometry, Graphs, and Networks
📅 21st November – 22nd November 2024 📍 Jersey City

Important dates:
- Registration deadline: November 14th 2024
Registration
Registration for this event is open until Nov 14th, 2024. Registration is free but required for attendance.
In addition, participants can optionally submit a poster for presentation during the poster session. Examples of research areas that are within scope for the poster session are described below under Subject Areas.
To register for participation or to submit a poster, please fill out this form.
Subject Areas
The following is a summary of LoG’s focus, which is not exhaustive. If you doubt that your paper fits the venue, feel free to contact logmeetupnyc@gmail.com!
- Expressive graph neural networks
- GNN architectures (e.g., transformers, new positional encodings, etc.)
- Statistical theory on graphs
- Causal inference and causal discovery (e.g., structural causal models, causal graphical models, etc.)
- Geometry processing and optimization
- Robustness and adversarial attacks on graphs
- Combinatorial optimization and graph algorithms
- Graph kernels
- Graph signal processing and spectral methods
- Graph generative models
- Scalable graph learning models and methods
- Graphs for recommender systems
- Knowledge graphs
- Neural manifold
- Self-supervised learning on graphs
- Structured probabilistc inference
- Graph/Geometric ML (e.g., for health applications, security, computer vision, etc.)
- Graph/Geometric ML infrastructures (e.g., datasets, benchmarks, libraries, etc.)
Schedule
The official event schedule is outlined below:
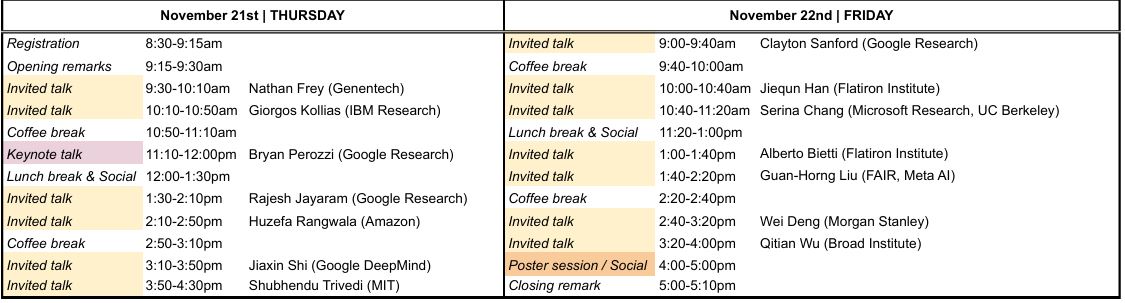
Download a PDF version of the schedule here.
Speakers

Thursday 9h30-10h10 Nathan Frey
Scalable Active Learning for Protein Design
Affiliation: Genentech
Abstract: We will discuss our approach and general considerations for implementing active learning and design of experiments to iteratively optimize proteins. Our framework is underpinned by robust data pipelines and algorithmic innovations, including novel strategies for building, pre-training, and fine-tuning foundation models for biology. I'll compare and contrast sequence- and structure-based protein design paradigms and discuss opportunities to think beyond these traditional mindsets.
Biography: Nathan Frey is a Principal Machine Learning Scientist and Group Leader at Prescient Design, Genentech. At Prescient, his group develops and applies machine learning methods to molecular discovery and design. Previously, Nathan was a Postdoctoral Associate at MIT working with the Lincoln Lab Supercomputing and AI groups. He was a National Defense Science & Engineering Graduate Fellow at the University of Pennsylvania, where he obtained his PhD in Materials Science & Engineering. During graduate school, Nathan was an affiliate scientist with the Materials Project at Berkeley Lab.

Thursday 10h10-10h50 Giorgos Kollias
Directed Graph Transfomers
Affiliation: IBM Research
Abstract: TBA
Biography: Dr. Georgios Kollias is a Research Staff Member (RSM) currently working on memory-augmented LLMs and graph NNs at IBM T.J. Watson Research Center, USA. He obtained the BSc in Physics in 2000 and the MSc in Computational Science in 2002 from the University of Athens, Greece, and the PhD in Computer Science from the University of Patras, Greece, in 2009. He then moved to Purdue University, USA and worked as a Postdoctoral Research Associate in the Computer Science Department and the Center for Science of Information till April 2013. Next, he joined IBM Research, USA also holding a research position at IBM Zurich Research Lab, Switzerland (August 2014 - April 2015). His research interests span the areas of graph mining and analytics, parallel, distributed and high-performance computing, numerical linear algebra and matrix computations, big data analytics, problem solving environments, quantum computations, nonlinear dynamical systems and machine learning applications.

Thursday 11h10-12h00 Bryan Perozzi
Challenges and Solutions in Applying Graph Neural Networks at Google
Affiliation: Google Research
Abstract: In this talk I'll look back at some challenges and lessons learned over the past seven years of applying GNNs to a wide variety of problems at Google. Interestingly, although the field has made substantial progress, many (if not all) of these challenges still remain.
Biography: Bryan Perozzi is a Research Scientist in Google Research’s Algorithms and Optimization group, where he routinely analyzes some of the world’s largest (and perhaps most interesting) graphs. Bryan’s research focuses on developing techniques for learning expressive representations of relational data with neural networks. These scalable algorithms are useful for prediction tasks (classification/regression), pattern discovery, and anomaly detection in large networked data sets. # Bryan is an author of 40+ peer-reviewed papers at leading conferences in machine learning and data mining (such as NeurIPS, ICML, ICLR, KDD, and WWW). His doctoral work on learning network representations (DeepWalk) was awarded the prestigious SIGKDD Dissertation Award. Bryan received his Ph.D. in Computer Science from Stony Brook University in 2016, and his M.S. from the Johns Hopkins University in 2011.

Thursday 13h30-14h10 Rajesh Jayaram
Multi-Vector Representations and Embedding-Based Nearest Neighbor Search
Affiliation: Google Research
Abstract: Perhaps the biggest breakthrough in information retrieval in the 21st century has been the usage of embedding models. Given a dataset D which could consist of images, sentences, videos, or other modalities, these models map each datapoint in D to a vector in high-dimensional Euclidean space, such that similar datapoints (e.g. similar images) are mapped to similar vectors under the Euclidean distance. This transforms a non-mathematical similarity to a mathematical one, reducing information retrieval to Euclidean Nearest Neighbor Search, which has been studied extensively in theory for over three decades. While this paradigm has been extremely successful, the representation of data by a single vector has limitations. In particular, it must compress all aspects of the data into a single global representation. Unfortunately, the actual similarity between two data-points can depend on a complex interaction between multiple local features of the data (e.g. words, subregions of an image, subsections of a document, ect.). Beginning with the landmark ColBERT paper (Khattab and Zaharia, SIGIR 2020), this has been addressed by training models to produce multiple embeddings per data-point. To measure the similarity between two sets of vectors A,B, one uses the so-called Chamfer Distance, which is the average distance between each point in A and its nearest point in B. Multi-vector models currently achieve SOTA on many retrieval benchmarks, and are now an extremely popular area of research. However, the usage of the Chamfer Distance raises a host of totally unexplored algorithmic questions. Firstly, Chamfer Distance is more expensive to compute than Euclidean distance. Is it possible to compute it faster, even approximately? Furthermore, to be useful for information retrieval, we need fast nearest neighbor search algorithms for Chamfer, but do such algorithms exist? In this talk, we investigate these algorithmic questions, and explore how techniques from TCS can be applied to expand the possibilities of practical information retrieval.
Biography: Rajesh is a Research Scientist at Google Research NYC, where he is part of the Algorithms and Optimization Group. Previously, he completed his PhD at Carnegie Mellon University, where he was advised by Professor David P. Woodruff. His research focuses on algorithms for high-dimensional geometry, especially streaming, sketching, and sublinear algorithms. He is the winner of two PODS best paper awards (2019, 2020) for his work on adversarial streaming algorithms and database theory, and has publications spanning a wide variety of other algorithmic domains, including property testing, randomized numerical linear algebra, nearest neighbor search, distributed algorithms, clustering, geometric streaming, and learning theory.

Thursday 14h10-14h50 Huzefa Rangwala
Leveraging Structured Knowledge for Generative AI Applications
Affiliation: Amazon
Abstract:
Biography: At AWS AI/ML, Huzefa Rangwala leads a team of scientists and engineers, revolutionizing AWS services through advancements in graph machine learning, reinforcement learning, AutoML, low-code/no-code generative AI, and personalized AI solutions. His passion extends to transforming analytical sciences with the power of generative AI. He is a Professor of Computer Science and the Lawrence Cranberg Faculty Fellow at George Mason University, where he also served as interim Chair from 2019-2020. He is the recipient of the National Science Foundation (NSF) Career Award, the 2014 University-wide Teaching Award, Emerging Researcher/Creator/Scholar Award, the 2018 Undergraduate Research Mentor Award. In 2022, Huzefa co-chaired the ACM SIGKDD conference in Washington, DC. His research interests include structured learning, federated learning, and ML fairness inter-twinned with applying ML to problems in biology, biomedical engineering, and learning sciences

Thursday 15h10-15h50 Jiaxin Shi
Discrete generative modeling with masked diffusions
Affiliation: Google DeepMind
Abstract: Modern generative AI has developed along two distinct paths: autoregressive models for discrete data (such as text) and diffusion models for continuous data (like images). Bridging this divide by adapting diffusion models to handle discrete data represents a compelling avenue for unifying these disparate approaches. However, existing work in this area has been hindered by unnecessarily complex model formulations and unclear relationships between different perspectives, leading to suboptimal parameterization, training objectives, and ad hoc adjustments to counteract these issues. In this talk, I will introduce masked diffusion models, a simple and general framework that unlock the full potential of diffusion models for discrete data. We show that the continuous-time variational objective of such models is a simple weighted integral of cross-entropy losses. Our framework also enables training generalized masked diffusion models with state-dependent masking schedules. When evaluated by perplexity, our models trained on OpenWebText surpass prior diffusion language models at GPT-2 scale and demonstrate superior performance on 4 out of 5 zero-shot language modeling tasks. Furthermore, our models vastly outperform previous discrete diffusion models on pixel-level image modeling, achieving 2.75 (CIFAR-10) and 3.40 (ImageNet 64×64) bits per dimension that are better than autoregressive models of similar sizes.
Biography: Jiaxin Shi is a research scientist at Google DeepMind. Previously, he was a postdoctoral researcher at Stanford and Microsoft Research New England. He obtained his Ph.D. from Tsinghua University. His research interests broadly involve probabilistic and algorithmic models for learning as well as the interface between them. Jiaxin served as an area chair for NeurIPS and AISTATS. He is a recipient of Microsoft Research PhD fellowship. His first-author paper was recognized by a NeurIPS 2022 outstanding paper award.

Thursday 15h50-16h30 Shubhendu Trivedi
Efficient Approximately Equivariant Neural Networks via Symmetry-Based Structured Matrices
Affiliation:
Abstract: There has been much recent interest in designing symmetry-aware neural networks exhibiting relaxed equivariance. Such NNs aim to interpolate between being exactly equivariant and being fully flexible, affording consistent performance benefits. This talk will present a method to construct approximately equivariant networks that perform roughly at par with the state of the art equivariant networks, but usually operate with one or two orders of magnitude fewer parameters. The construction is based on a novel generalization of classical low-displacement rank theory, which works only for the cyclic group, to general discrete groups and their homogeneous spaces. The generalization allows the design of special structured matrices specific to a group of interest that allows for generalizing equivariant networks, and provide a recipe for controlling approximation error. Ongoing work on efficient tensorizations and GPU implementations will also be discussed.
Biography: Shubhendu Trivedi works on the theoretical and applied aspects of machine learning. For the past few years his research has focused on developing rigorous theoretical and engineering tools for data-efficient machine learning (e.g. via equivariant and geometric deep learning) and enabling their safe and reliable deployment in real-world applications and decision-making pipelines (e.g. via conformal prediction and provable uncertainty quantification). In research contexts, he has spent time at the MIT CSAIL, Brown University, Fermilab, University of Chicago, TTI-Chicago, WPI, and for industrial work he has been involved with data science consulting in the pharmaceuticals, airlines, and FMCG sectors, and has also spent time at ZS, NEC, United, amongst others. In addition to the above work, he has co-founded a startup in semiconductors and serves on the boards of multiple startups, the most recent of which include Reexpress (LLMs), Brainwell Health (Imaging), and Spark Neuro (EEG).

Thursday 16h30-16h45 Ali Parviz
NLSD-GNN: The Power of Non-linearity for Signed-Directed Graph Learning
Affiliation: MILA
Abstract: While signed-directed graphs have been studied using linear Laplacians in the design of graph neural networks, relatively little research has focused on developing non-linear Laplacian operators for such networks. We introduce a new non-linear Laplacian operator specific to signed and directed networks (NLSD). This non-linear operator extends the concepts of the signed Laplacian for signed graphs and the standard Laplacian for directed graphs. The NLSD calculates node-specific potentials based on features, leveraging message-passing techniques only across edges where potential discrepancies align with the edge's direction. Utilizing this novel operator, we propose an efficient spectral GNN framework (NLSD-GNN). We conducted comprehensive evaluations focusing on node classification and link prediction, examining scenarios involving signed, directional, or both types of information. Our findings reveal that this spectral GNN framework not only integrates signed and directional data effectively but also achieves superior performance across diverse datasets.
Biography:

Thursday 16h45-17h00 Hannah Lawrence
Equivariant Frames and the Impossibility of Continuous Canonicalization
Affiliation: MIT
Abstract: Canonicalization provides an architecture-agnostic method for enforcing symmetries, with generalizations such as frame-averaging recently gaining prominence as a lightweight and flexible alternative to equivariant architectures. In this work, we elucidate an unfortunate property of canonicalization: for commonly used groups, there is no efficiently computable choice of frame that preserves continuity of the underlying neural network. We address this fundamental robustness problem by constructing weighted frames, which provably preserve continuity while remaining efficient to implement.
Biography:

Friday 09h00-09h40 Clayton Sanford
Understanding Transformer Reasoning Capabilities via Graph Algorithms
Affiliation: Google Research
Abstract: Which transformer scaling regimes are able to solve different classes of algorithmic problems? Despite the tremendous empirical advances of transformer-based neural networks, a theoretical understanding of their algorithmic reasoning capabilities in realistic parameter regimes is lacking. We investigate this question in terms of the network's depth, width, and number of extra tokens for algorithm execution. We introduce a novel representational hierarchy that separates graph algorithmic tasks, such as connectivity and shortest path, into equivalence classes solvable by transformers in different realistic parameter scaling regimes. We prove that logarithmic depth is necessary and sufficient for tasks like graph connectivity, while single-layer transformers with small embedding dimensions can solve contextual retrieval tasks. We supplement our theoretical analysis with empirical evidence using the GraphQA benchmark. These results show that transformers excel at many graph reasoning tasks, even outperforming specialized graph neural networks.
Biography: Clayton Sanford is a research scientist at Google and a recent PhD graduate from Columbia University. In his PhD, he studied machine learning and theoretical computer science under the supervision of Professors Daniel Hsu and Rocco Servedio. His research focuses on developing a more rigorous understanding of neural network expressivity and generalization, with a particular focus on modern language models.

Friday 10h00-10h40 Jiequn Han
Provable Posterior Sampling with Score-Based Diffusion via Tilted Transport
Affiliation: Flatiron Institute
Abstract: Score-based diffusion models have significantly advanced the generation of high-dimensional data across various domains by learning a denoising oracle (or score) from datasets. From a Bayesian perspective, these models offer a realistic approach to modeling data priors and facilitate solving inverse problems through posterior sampling. Despite the development of numerous heuristic methods for this purpose, most lack the quantitative guarantees required for scientific applications. This talk introduces a technique called tilted transport, based on the analysis of SDEs for data generation. We demonstrate that the denoising oracle enables an exact transformation of the original posterior sampling problem into a new `boosted' posterior that is provably easier to sample from. We quantify the conditions under which this boosted posterior is strongly log-concave, highlighting the dependencies on the condition number of the measurement matrix and the signal-to-noise ratio.
Biography: Jiequn Han is a Research Scientist in the Center for Computational Mathematics, Flatiron Institute, Simons Foundation. He conducts research on machine learning for science, drawing inspiration from various scientific disciplines and focusing on solving high-dimensional problems in scientific computing, primarily those related to PDEs.

Friday 10h40-11h20 Serina Chang
Inferring and Simulating Human Networks with Machine Learning
Affiliation: Microsoft Research, UC Berkeley
Abstract: Understanding human networks is crucial for high-stakes decision making. For example, pandemic response requires understanding how disease spreads through contact networks and how individuals alter their behavior in response to policies and disease. However, human networks are often difficult to observe (e.g., for privacy reasons or data collection constraints). In this talk, I'll discuss two approaches to addressing this challenge: (1) inferring networks from novel data sources, (2) simulating networks and behaviors with LLMs. In the first part, I'll discuss our work to infer fine-grained mobility networks from aggregated location data, which enabled us to model the spread of COVID-19 in cities across the US and inform public health decision-making. In the second part, I'll discuss our recent work on generating social networks with LLMs, showing that, while these models can capture structural characteristics of real-world networks, they substantially overestimate political homophily.
Biography: Serina Chang is an incoming Assistant Professor at UC Berkeley, with a joint appointment in EECS and Computational Precision Health. She recently completed her PhD in CS at Stanford University, advised by Jure Leskovec and Johan Ugander, and is currently a postdoc at Microsoft Research NYC. Her research develops AI and graph methods to study human behavior, improve public health, and guide data-driven policymaking. Her work is recognized by a KDD Best Paper Award, NSF Graduate Research Fellowship, Meta PhD Fellowship, EECS Rising Stars, Rising Stars in Data Science, and Cornell Future Faculty Symposium, and has received coverage from over 650 news outlets, including The New York Times and The Washington Post.

Friday 13h00-13h40 Alberto Bietti
Associative memories as a building block in Transformers
Affiliation: Flatiron Institute
Abstract: Large language models based on transformers have achieved great empirical successes. However, as they are deployed more widely, there is a growing need to better understand their internal mechanisms in order to make them more reliable. These models appear to store vast amounts of knowledge from their training data, and to adapt quickly to new information provided in their context or prompt. Through toy tasks for reasoning and factual recall, we highlight the role of weight matrices as associative memories, and provide theoretical results on how gradients enable their learning during training, and how over-parameterization affects their storage capacity.
Biography: Alberto Bietti is a research scientist at the Center for Computational Mathematics (CCM) in the Flatiron Institute of the Simons Foundation. He received his Ph.D. in applied mathematics from Inria Centre at the University Grenoble Alpes in 2019 and was a Faculty Fellow at the NYU Center for Data Science from 2020 to 2022. He also spent time at Inria Paris Centre, Microsoft Research, and Meta AI. His research focuses on the foundations of deep learning.

Friday 13h40-14h20 Guan-Horng Liu
Learning Scalable Diffusion Models using Optimality and Constraint Structures
Affiliation: Meta
Abstract: Generative AI has made remarkable strides in recent years, largely propelled by the development of diffusion models. However, the successes of diffusion models largely rely on the structure of the data-to-noise diffusion processes, posing challenges for more general distribution matching problems beyond generative modeling. In this talk, I will present some of my recent works on learning scalable diffusion models for general distribution matching. Specifically, I will demonstrate how optimality and constraint structures can serve as a principled way for algorithmic design, generalizing those used in training denoising diffusion models, and achieve superior performance in image restoration, unsupervised image translation, and watermarked image generation.
Biography: Guan-Horng Liu is a Research Scientist at Fundamental AI Research (FAIR) at Meta in NY. He holds a Ph.D. in Machine Learning from Georgia Tech, where he was supervised by Evangelos Theodorou, and a M.S. in Robotics from Carnegie Mellon University. His research lies in fundamental algorithms for learning diffusion models with optimality structures, with the goals of enhancing theoretical understanding and developing large-scale algorithms for novel applications. His papers have been awarded with spotlight and oral presentations in ICLR, ICML, NeurIPS. He was supported by the Graduate Fellowship from the School of Aerospace Engineering at Georgia Tech.

Friday 14h40-15h20 Wei Deng
Variational Schrödinger Diffusion Models and Beyond
Affiliation: Morgan Stanley
Abstract: Schrödinger bridge (SB) methods are crucial for optimizing transportation plans in diffusion models, but they are often costly and complex due to intractable forward score functions. To address these issues and improve scalability, we introduce the Variational Schrödinger Diffusion Model (VSDM) and its momentum extensions. VSDM uses variational inference to simplify the forward score functions, creating a simulation-free forward process for training backward scores. This model leverages stochastic approximation to train its variational scores, balancing accuracy and computational budget. To further enhance transport and scalability, we explore momentum accelerations, which simplify the denoising process and reduce tuning costs. Empirically, both models demonstrate remarkable performance in anisotropic optimal transport and image generation, proving more scalable than classical SB methods without requiring warm-up training. Additionally, the conditional versions show exceptional performance in time series prediction.
Biography: Wei Deng is a machine learning researcher at Morgan Stanley, NY. He completed his Ph.D. from Purdue University in 2021. His research focuses on Monte Carlo methods, Diffusion Models, and State Space Models. His objective is to develop more scalable and reliable probabilistic methods for solving machine learning applications in Bayesian inference, generative models, and time series. His works are mainly published in machine learning conferences, such as NeurIPS, ICML, ICLR, UAI, AISTATS and journals such as Statistics and Computing and Journal of Computational and Graphical Statistics.

Friday 15h20-16h00 Qitian Wu
Towards Graph Transformers at Scale
Affiliation: Broad Institute of MIT and Harvard
Abstract: Representation learning over graph-structured data is a long-standing fundamental challenge in learning on graphs. Recent advances have demonstrated the promise of Transformer-style models that leverage global all-pair attentions for graph representation learning. However, the quadratic complexity and complicated architectures play as two computational bottlenecks for Transformers in large-scale data, and beyond accuracy criteria, it lacks principled guidance for interpretable model designs. This talk will introduce three recent works that present a scalable graph Transformer with linear complexity (NodeFormer), a simplified Transformer with single-layer global propagation (SGFormer), and an energy-constrained diffusion framework that reveals the inherent connections between graph neural networks and Transformers (DIFFormer). Future directions will be discussed at the end of this talk.
Biography: Qitian Wu is a postdoctoral fellow at Broad Institute of MIT and Harvard. Prior to this, he achieved PhD in Computer Science at Shanghai Jiao Tong University. His research interest focuses on machine learning with complex structured data. His recent works endeavor to develop efficient foundational backbones for representing large-scale graph data and provably generalizable learning algorithms for handling distribution shifts. He also seek to apply this methodology to address the pressing problems in recommender systems and biomedical science. He is the recipient of Eric and Wendy Schmidt Center Fellowship, Microsoft Research PhD Fellowship, and Baidu PhD Scholarship.
Spotlight Talks
Organizers
Sponsors
The Venue
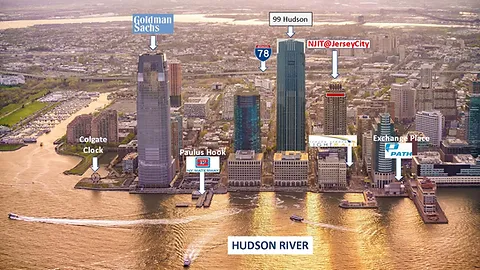
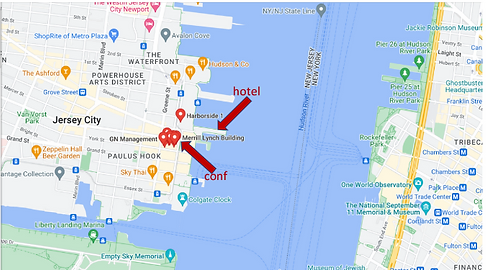
The New York LoG Meet Up will take place in the “NJIT@Jersey City Campus”. The NJIT@JerseyCity is located on the 36th floor of 101 Hudson Street on the Waterfront of Jersey City. The building occupies an entire block bordered by Montgomery Street, Hudson Street, York Street, and Greene Street. “Directions from Google Maps”
Public Transportation
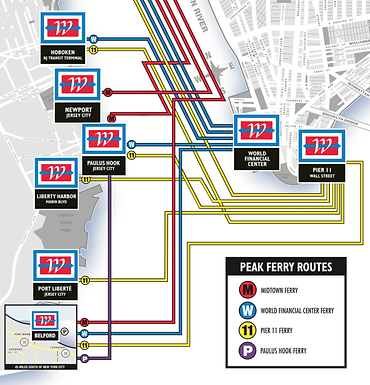
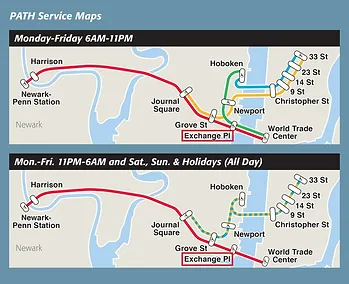
101 Hudson Street is located across the street from Exchange Place station for the PATH and the Hudson-Bergen Light Rail. The PATH at Exchange Place is served by the Newark-World Trade Center and Hoboken-World Trade Center lines. The light rail at Exchange Place is serviced by two routes: the Tonnelle Avenue - West Side Avenue and the Hoboken Terminal - 8th Street service routes. The NY Waterway Paulus Hook Terminal is located at the end of Sussex Street at the Hudson River Waterway three blocks away (10 minutes walking distance) from 101 Hudson Street. There are three ferry routes from NYC: Pier 11/Wall Street., Downtown/Brookfield Place, and Midtown/W. 39th Street.
All building visitors must check in with the security and receive a temporary pass. Walk to the middle elevator bank to access the 36th floor. Exit the elevator and see signs for NJIT@JerseyCity. Turn right to the double glass door entrance to NJIT@JerseyCity (Suite 3610).
Parking and Building Entry
To access the parking garage continue along Hudson Street to York Street; turn right onto York Street and continue towards Greene Street or approach the parking garage by driving along Greene Street and taking a left onto York Street. The entrance to the garage will be on the left.
The entrance to the parking garage is the last "garage door" entrance before the intersection of York and Greene Streets: Google Image of Parking Deck
To enter the parking garage push the button on the ticket dispensing machine; retrieve (and keep!) the ticket to release the arm of the gate to continue driving up the steep ramp to Level 2. Parking is available on Levels 2 - 5. Keep the parking ticket to process payment and to be able to exit the parking deck. Once parked, see signs for the elevators to the lobby.
Exit the garage elevator onto the rear lobby of 101 Hudson Street. Turn right to walk down the corridor towards the front of the building.
All building visitors must check in with the security and receive a temporary pass. Walk to the middle elevator bank to access the 36th floor. Exit the elevator and see signs for NJIT@JerseyCity. Turn right to the double glass door entrance to NJIT@JerseyCity (Suite 3610).